-
Notifications
You must be signed in to change notification settings - Fork 1
Home
Welcome to MetaFX wiki page!
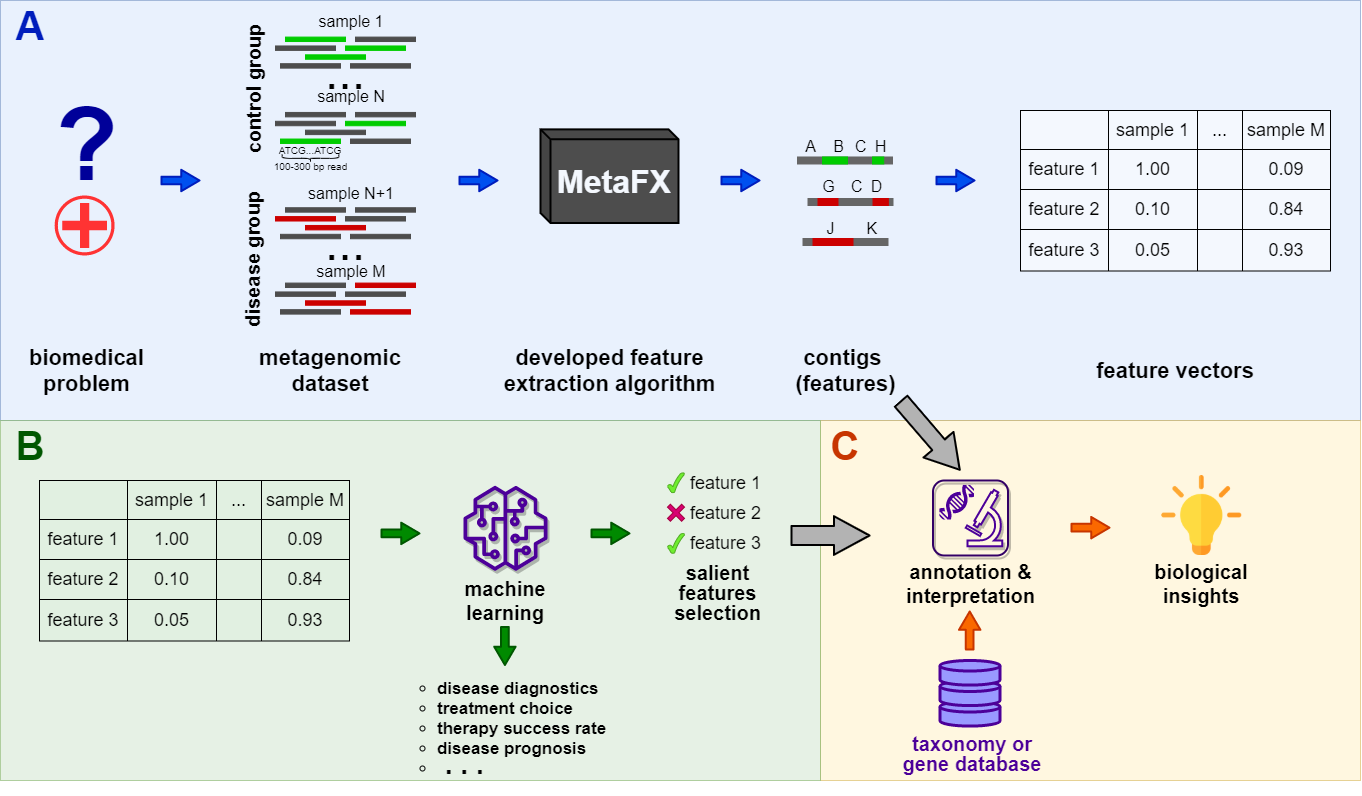
MetaFX (METAgenomic Feature eXtraction) is an open-source library for feature extraction from whole-genome metagenome sequencing data and classification of groups of samples.
The idea behind MetaFX is to introduce the feature extraction algorithm specific for metagenomics short reads data. It is capable of processing hundreds of samples 1-10 Gb each. The distinct property of suggest approach is the construction of meaningful features, which can not only be used to train classification model, but also can be further annotated and biologically interpreted.
To run MetaFX, one need to clone repo with all binaries and add them to PATH.
git clone https://github.com/ivartb/metafx_new
cd metafx_new
export PATH=bin:$PATHRequirements:
- JRE 1.8 or higher
- python3
- python libraries listed in
requirements.txtfile. Can be installed using pip
python -m pip install --upgrade pip
pip install -r requirements.txt- If you want to use
metafx metaspadespipeline, you will also need SPAdes software. Please follow their installation instructions (not recommended for first-time use).
Scripts have been tested under Ubuntu 18.04 LTS and Ubuntu 20.04 LTS, but should generally work on Linux/MacOS.
To run MetaFX use the following syntax:
metafx <pipeline> [<Launch options>] [<Input parameters>]To view the list of supported pipelines run metafx -h or metafx --help.
To view help for launch options and input parameters for selected pipeline run metafx <pipeline> -h or metafx <pipeline> --help.
By running MetaFX a working directory is created (by default ./workDir/).
All intermediate files and final results are saved there.
MetaFX is a toolbox with a lot of modules divided into three groups:
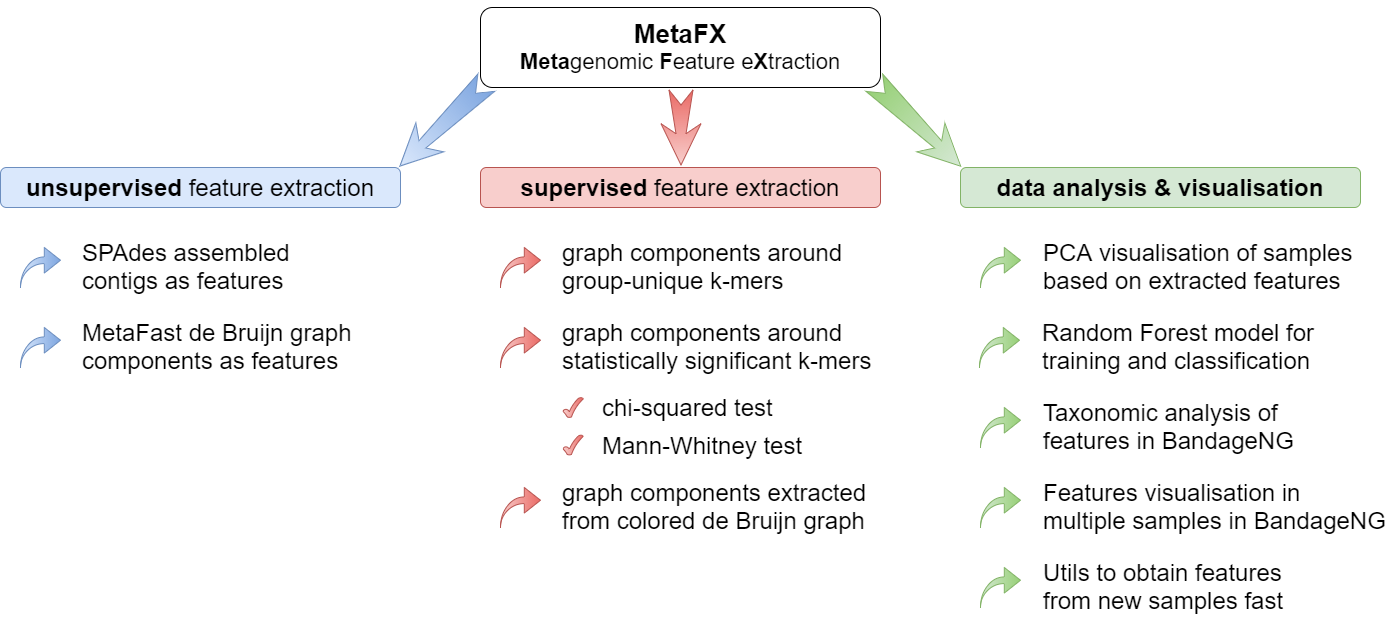
There are pipelines aimed to extract features from metagenomic dataset without any prior knowledge about samples and their relations. Algorithms perform (pseudo-)assembly of samples separately and construct the de Bruijn graph common for all samples. Further, graph components are extracted as features and feature table is constructed.
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
There are pipelines aimed to extract group-relevant features based on metadata about samples such as diagnosis, treatment, biochemical results, etc. Dataset is split into groups of samples based on provided metadata information and group-specific features are constructed based on de Bruijn graphs. The resulting features are combined into feature table.
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
There are pipelines for analysis of the feature extraction results. Methods for samples similarity visualisation and training machine learning models are implemented. Classification models can be trained to predict samples' properties based on extracted features and to efficiently process new samples from the same environment.
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO TODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODOTODO
Minimal example is provided in the README page.
More sophisticated example is presented as a jupyter notebook with links to input and output data, all commands and commentaries.